Multi-modal Graph Learning over UMLS Knowledge Graphs
Abstract
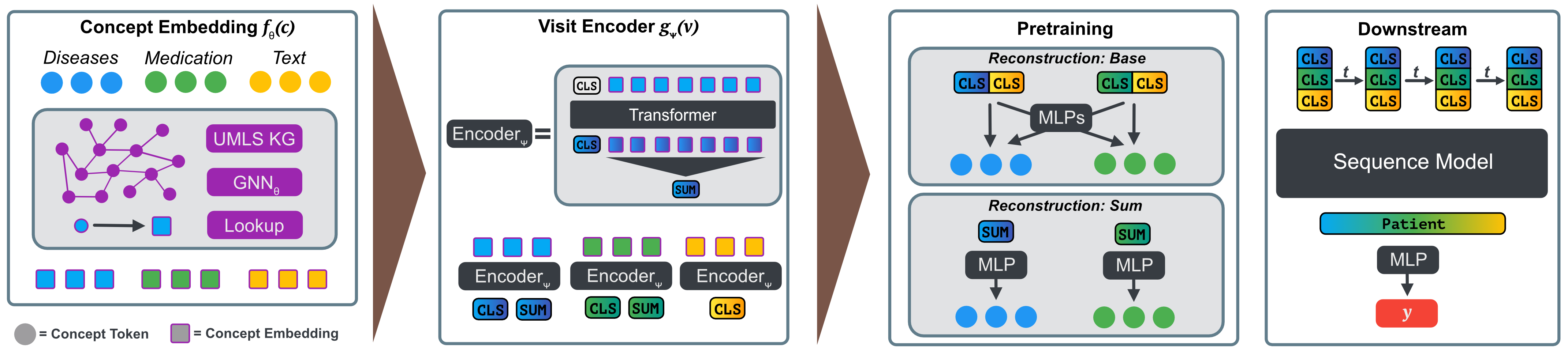
Clinicians are increasingly looking towards machine learning to gain insights about patient evolutions. We propose a novel approach named Multi-Modal UMLS Graph Learning (MMUGL) for learning meaningful representations of medical concepts using graph neural networks over knowledge graphs based on the unified medical language system. These representations are aggregated to represent entire patient visits and then fed into a sequence model to perform predictions at the granularity of multiple hospital visits of a patient. We improve performance by incorporating prior medical knowledge and considering multiple modalities. We compare our method to existing architectures proposed to learn representations at different granularities on the MIMIC-III dataset and show that our approach outperforms these methods. The results demonstrate the significance of multi-modal medical concept representations based on prior medical knowledge.
TL;DR
In this work, published in the Proceedings of Machine Learning (PMLR) and presented at the Machine Learning for Health (ML4H) conference 2023 in New Orleans, we have extracted a large medical knowledge graph from the Unified Medical Language System (UMLS) and trained it using electronic health records (EHR) of patients.
We show increased predictive performance for a range of clinical prediction tasks such as the onset of heart failure, medication recommendation, and readmission. The performance improvements can be attributed to the inclusion of rich prior knowledge extracted from UMLS, the training on rich multi-modal data (structured and unstructured EHR), and the choice of pretraining.
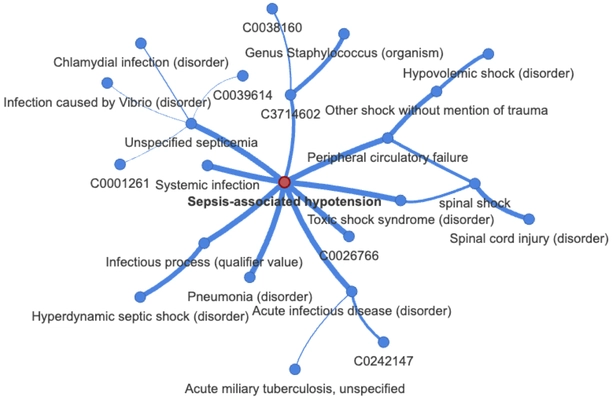
Beyond performance improvements we also ascertain the interpretability of our model by using GNNExplainer (Ying et al., 2019) to extract subgraphs relevant for the embedding of specific concepts (and as such predictions making use of said concept). The figure below shows the subgraph relevant for the concept of Septic Shock (Sepsis-associated hypotension, UMLS C0036983) extracted using GNNExplainer. Edge thickness corresponds to the edge weight computed by GNNExplainer and the explanation is a reduction of 9932 nodes in a 3-hop neighborhood to 23 related nodes.
Citation
| |
